Key Points:
- Researchers at the University of Chicago developed a breakthrough system to map gene expression throughout the entire mouse body, rather than focusing on a single organ.
- The team successfully identified and mapped approximately 75% of all known cell types within a mouse cross-section using high-resolution spatial transcriptomics.
- A new machine learning model allows scientists to virtually label organs and cells, replacing slow, expensive manual staining methods with a cheaper, faster digital process.
- This technology lays the foundation for a “virtual mouse,” a digital model that could eventually replace thousands of live animals in drug discovery and disease research.
For decades, biological research hit a frustrating wall. Scientists could easily study gene expression within a single organ or a tiny slice of tissue, but they struggled to see how diseases impacted the whole body at once. A research team at the University of Chicago’s Pritzker School of Molecular Engineering just broke that barrier. Led by Associate Professor Nicolas Chevrier and staff scientist Maggie Clevenger, the group developed an interdisciplinary system that maps molecular and cellular processes across an entire mouse body.
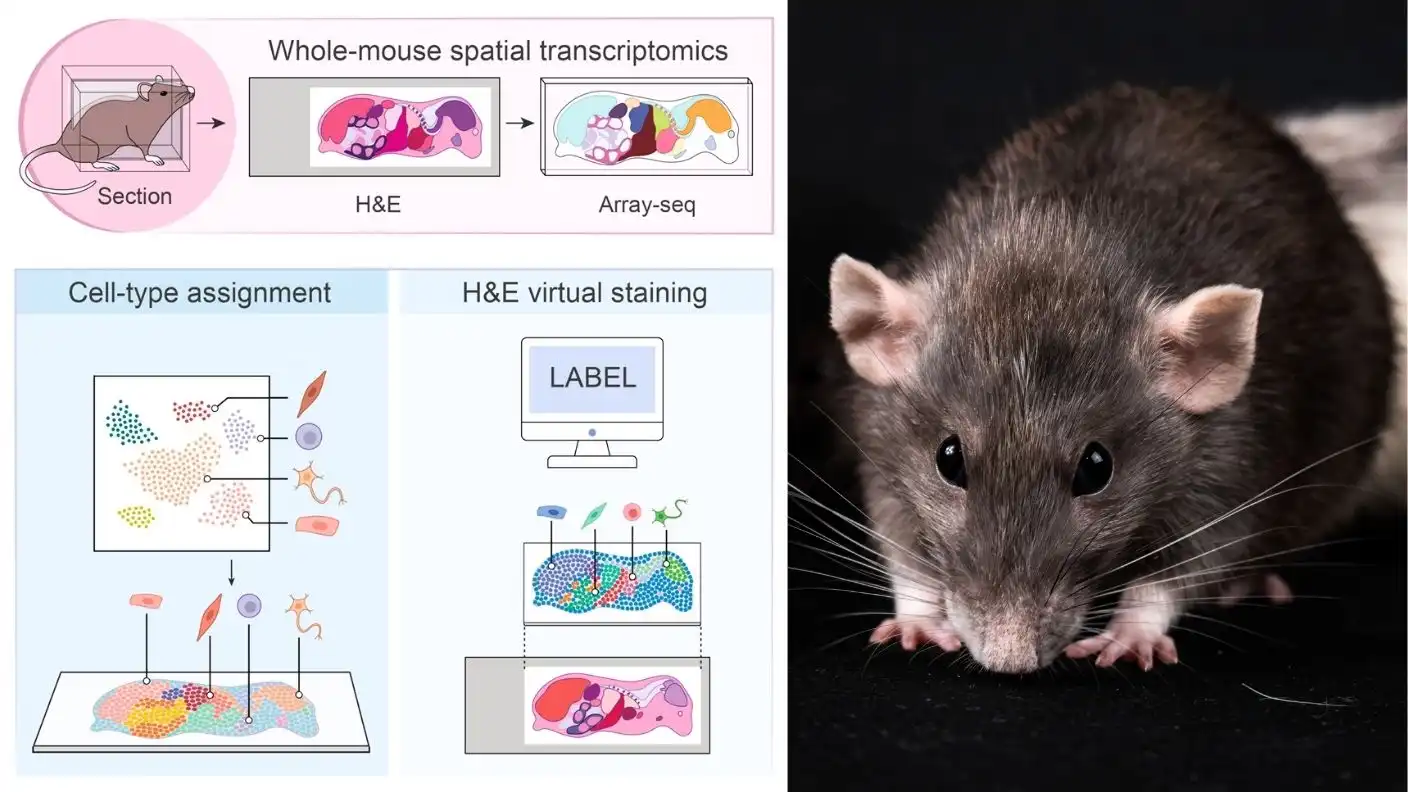
The team combined spatial transcriptomics—a method that uses high-resolution microscopy and genetic sequencing—with powerful new computational tools. By optimizing how they prepare delicate biological specimens, the researchers mapped every major organ and roughly 75% of all known cell types in the mouse. This achievement represents a massive leap forward for both basic science and the pharmaceutical industry, providing a toolkit that was previously considered unimaginable.
To make this possible, the team relied on a technique they developed in 2025 called Array-seq. This method uses DNA microarrays with custom probes to analyze tissue samples. However, slicing an entire mouse body into sections as thin as a single cell without destroying the fragile RNA inside proved incredibly difficult. Working with Professor Tadafumi Kawamoto at Tsurumi University in Japan, the team perfected a method to freeze and slice the specimen while preserving it intact for analysis.
After capturing the genetic data, the researchers turned to artificial intelligence to make sense of the overwhelming amount of information. They collaborated with Ashwini Patil from Combinatics to build a computational model that annotates cellular data across the entire mouse. They also recruited AI expert Professor Feng Bao from Fudan University to train a machine learning model. This model can identify and label different cell types and tissues on standard slides stained with hematoxylin and eosin, a routine clinical stain.
Doing this work manually would cost a fortune and take years, as scientists would need to label every cell type with specific, expensive antibodies. “We trained an AI model to do this, so now we can do it virtually and very cheaply,” Professor Chevrier explained. By avoiding the manual labor involved in traditional staining reagents, the team significantly reduced the complexity of whole-body mapping.
The researchers proved the power of their new system by studying sepsis, a dangerous, body-wide immune response to infection. For the first time, the team quantified exactly how systemic inflammation hits every major organ and cell type simultaneously. This level of detail offers insights that single-organ studies simply miss. It allows scientists to see how a new drug might trigger unexpected side effects in an organ they were not even monitoring, potentially saving millions in failed drug trials.
The team now looks toward the ultimate goal: building a “virtual mouse.” Rather than analyzing a single thin slice of an organism, they want to model the entire body in three dimensions. If successful, this digital mouse would allow researchers to test therapies and simulate complex diseases without needing to conduct experiments on thousands of live laboratory animals. This shift would represent a $5,000 to $10,000 reduction in costs for early-stage testing and a massive improvement in ethical research practices.
As the team continues to refine these computational tools, they hope to set a new standard for biological research. By mapping the molecular landscape of model organisms at this scale, they provide the scientific community with a clearer picture of how life functions as a whole system. This work, published in the journal Cell, marks a turning point in how we study, diagnose, and treat complex diseases in the 21st century.
Source: Cell (2026).