Key Points:
- When a cell activates a gene, it physically twists and unwinds the DNA strand, sending ripples that affect surrounding genes.
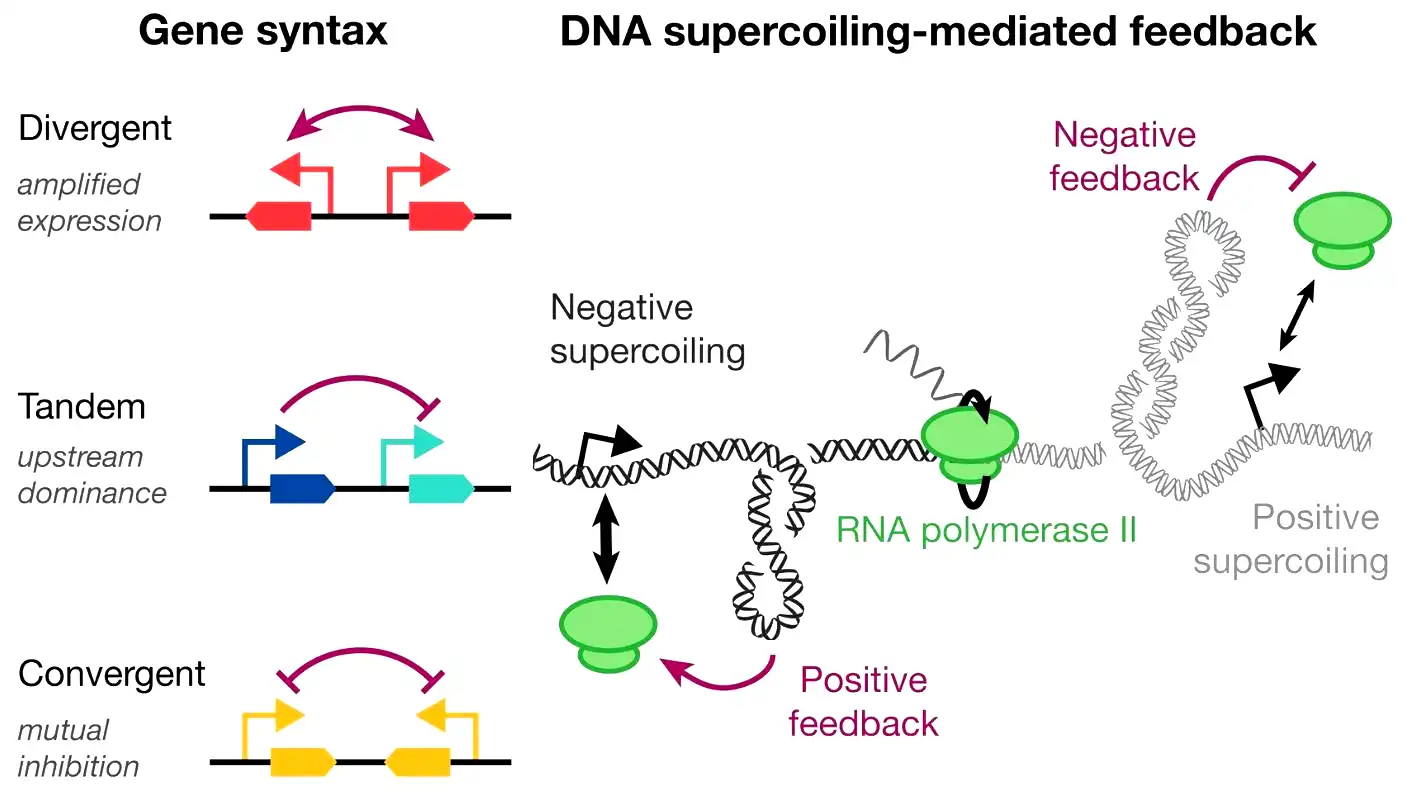
- Placing genes in a “divergent” layout facing away from each other significantly boosts the output of both genes.
- The physical ripples in DNA alter gene expression by up to 25-fold and extend up to 2,000 base pairs away.
- Scientists successfully used this discovery to physically rearrange DNA and produce much higher amounts of a yellow fever antibody.
When a cell turns on a gene, it sends out a physical ripple along the DNA strand. A recent study by MIT researchers, published in the journal Science, reveals that these ripples physically alter DNA structure to either boost or block neighboring genes.
To copy a gene into messenger RNA, an enzyme called RNA polymerase must unwind the double-stranded DNA helix. This unwinding loosens the DNA behind the active gene and tightens the DNA in front of it. Because the upstream DNA becomes looser, enzymes can more easily attach and activate nearby genes. Meanwhile, the tightened downstream DNA blocks enzymes and turns down gene activity.
The research team calls this physical arrangement “gene syntax.” Katie Galloway, an assistant professor of chemical engineering at MIT and the senior author of the study, explained that syntax lets scientists coordinate gene activity in entirely new ways. Before this discovery, scientists mostly built circuits using chemical triggers. Now, they can design the physical layout of DNA to achieve the exact results they want and to support dynamic changes within the cell.
Back in 2022, Galloway and lead author Christopher Johnstone used computer models to predict how these physical changes alter gene expression. They looked at 3 specific layouts: tandem, divergent, and convergent. In a tandem layout, one gene sits right behind another. A divergent layout places genes so they face away from each other, while a convergent setup points them toward each other. Most synthetic gene circuits in the past simply relied on a tandem arrangement.
To see if real cells matched their computer models, the team engineered these 3 different setups into human cell lines and human induced pluripotent stem cells. They wanted to observe the physical interactions firsthand. They discovered that their original predictions were highly accurate. The divergent circuits significantly amplified the output of both genes. In contrast, the tandem circuits suppressed the downstream gene the moment the upstream gene turned on.
These structural changes drastically impacted how the cells worked. The researchers recorded up to a 25-fold increase or decrease in gene expression. Furthermore, these physical ripple effects extended across distances of up to 2,000 base pairs between genes.
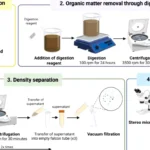
To map these changes, the team used a high-resolution tool called Region Capture Micro-C. They saw exactly what happens to the strand. The DNA downstream from an active gene twisted tightly into structures called plectonemes. These structures resemble the messy tangles you see on an old twisted telephone cord, making it nearly impossible for RNA polymerase to bind to the DNA.
To make this possible, the MIT researchers teamed up with scientists from Leiden University Medical Center. Together, they created a new system called STRAIGHT-IN Dual. Published in the journal Nature Biomedical Engineering, this tool enables scientists to efficiently insert 2 separate genes into the same DNA strand to test these physical layouts.
The team has already proved how useful this discovery can be in medicine. They built synthetic circuits carrying the genes for 2 segments of a new yellow fever antibody. By placing the genes in a divergent layout, the engineered human cells successfully produced much larger quantities of the antibody.
Moving forward, designers of synthetic gene circuits have a powerful new tool. They no longer have to rely only on biochemical activator molecules. They can just alter the physical layout of the DNA components.
Beyond antibodies, Galloway and her lab have already used this approach to improve the output of other synthetic gene circuits. They previously developed systems to deliver gene therapy or reprogram adult cells into completely different cell types. Knowing how to arrange the genes properly makes these therapies much more reliable.
Ultimately, this strategy opens the door to building complex, dynamic circuits inside living cells. Scientists can easily create toggle switches or pulse generators for any medical application requiring strict control of gene output. The researchers believe that understanding this physical syntax will pave the way for programming highly complex behaviors in future human health treatments.
Source: Science (2026)