Key Points:
- A major collaboration led by the Gladstone Institutes introduced a powerful DNA-editing system into 14 new bacterial species.
- The new approach uses retrons, a bacterial immune system that researchers have repurposed to efficiently edit genomes.
- The successful DNA editing spanned three major branches of the bacterial family tree, including species relevant to human health.
- This breakthrough will help scientists study dangerous diseases, develop sustainable materials, and fight drug-resistant infections.
Microbiologists have spent years chasing a massive scientific goal. They desperately want the ability to edit the genomes of different bacteria precisely. Having this specific technology would allow scientists to make incredible new inroads into studying complex diseases, developing highly sustainable materials, and fighting dangerous drug-resistant infections. Unfortunately, for many years, the most powerful tools available for bacterial genome editing only worked on Escherichia coli, commonly known as E. coli, which is the most common laboratory bacteria.
That limitation officially ended this week. A massive collaboration involving nine different laboratories, led primarily by scientists at Gladstone Institutes, successfully transferred a highly useful DNA editing system out of E. coli and into 14 brand new species of bacteria. This massive scientific leap spanned three major branches of the entire bacterial family tree. The researchers recently described their incredible approach in the journal Nature Biotechnology.
The scientists achieved this massive breakthrough by leveraging something called retrons. In nature, retrons act as a built-in immune system for bacteria. They function like a kind of viral alarm system, producing tiny strands of DNA to protect the cell.
Gladstone Investigator Seth Shipman, the senior author of the new study, explained how his lab completely repurposed these natural alarm systems. His team used the highly efficient DNA-making machinery found in retrons to synthesize tiny DNA strands. These factories produce the exact new strands of DNA needed for complex genome editing. Using these retrons, his team successfully created a tool that can efficiently modify DNA in bacteria, yeast, and even human cells. When used in bacteria, scientists call the transformed retron-derived editor a recombitron.
Shipman noted that his team has easily edited E. coli genomes using retrons for years. This ability substantially increased the overall pace of their fundamental biology research and advanced the development of molecular technology. However, scientists from the broader field constantly asked his team when a version of this specific technology would be ready to work in other bacterial species that actually matter for the environment, massive industrial processes, or critical human health.
To answer that question, the scientists wanted to aggressively probe whether they could engineer the retrons to work much more broadly. They carefully developed a highly diverse panel of 10 different retro-based editing systems. They actively collaborated with nine labs from around the world that specialized in working with a wide range of unusual bacteria.
Alejandro Gonstudy ‘s-Delgado, a scholar in Shipman’s lab and the study’s first author, explained the complex logistics of the massive experiment. The team designed all the physical molecular parts at the Gladstone facility. Then, they mailed the actual parts to the various global collaborators, who actually ran the spine experiments in their own labs. After finishing the work, each collaborator shipped the samples back to Gladstone, where Gonzalez-Delgado performed a massive, centralized, and highly in-depth analysis of the final results.
The massive team collected full data on exactly how well the retrons worked across 15 different bacterial species, with massive relevance across multiple scientific fields. Several of the tested species, including Klebsiella pneumoniae and Pseudomonas aeruginosa, are highly dangerous human pathogens that often develop severe antibiotic resistance. This specific success underscores the massive need for new research into exactly how to fight these deadly bacteria. Other tested species, such as Vibrio natriegens and Pseudomonas putida, are especially fast-growing and are commonly used in modern biotechnology to rapidly produce desired compounds, ranging from lifesaving medicines to complex fuels.
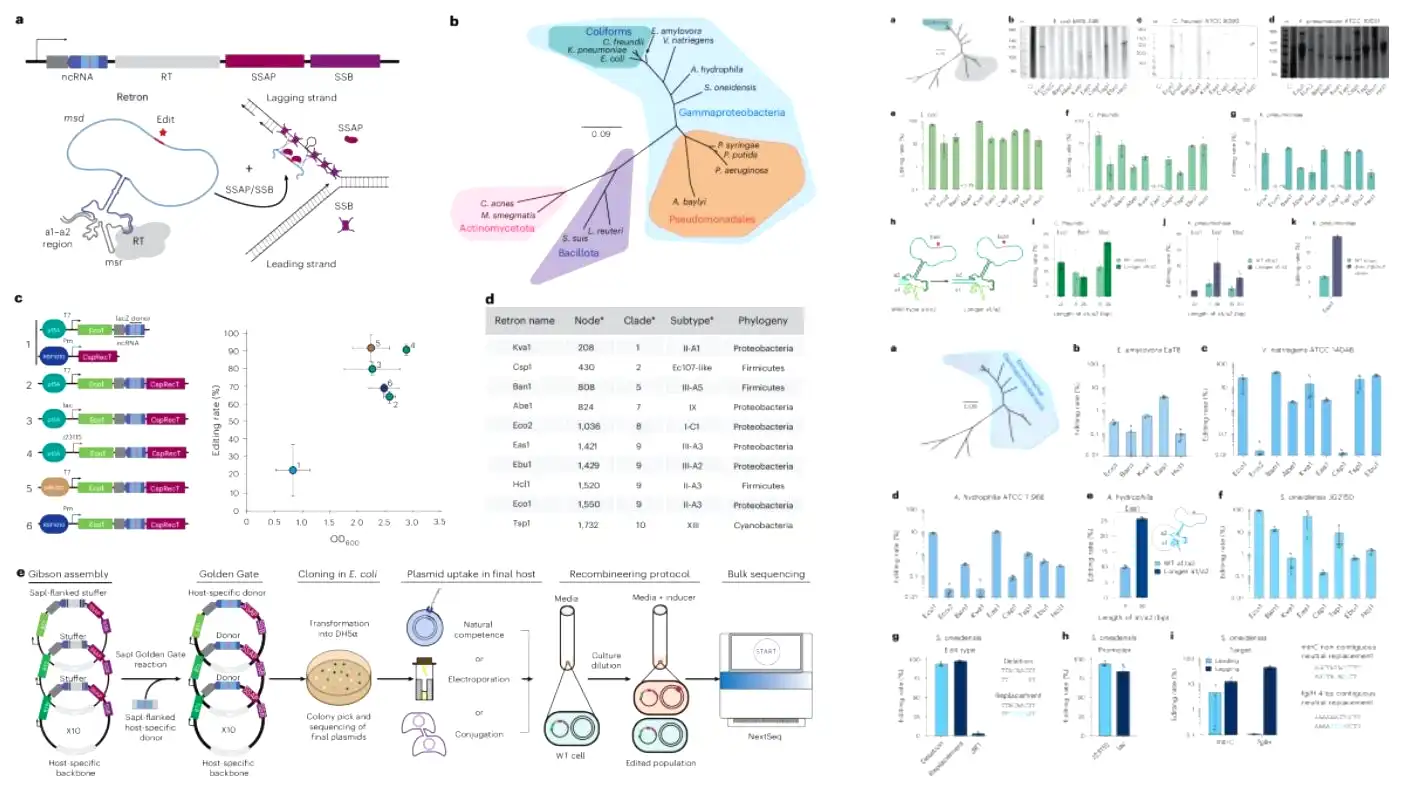
The incredible experiment was a total success. The retrons, packaged tightly with other specific proteins into genome-editing systems known as recombitrons, successfully worked in all 15 species tested. However, Gonzalez-Delgado noted that different retrons excelled in different specific species. He explained that each retron worked completely differently depending on the bacteria. This strange result reinforces why it remains so important to have many different retrons available, so scientists can easily choose the specific ones best suited to their favorite bacterial species.
The editing rates also varied wildly during the massive study. The editing rate simply measures the percentage of bacterial cells that actually carry the intended genetic change. Some species saw editing rates at just a tiny fraction of a percent, while other species saw massive success rates exceeding 90%. For those stubborn species with much lower editing rates, the team successfully demonstrated that slightly altering the retron’s physical structure or other components of the complex editing system could easily boost final editing rates.
This massive scientific success opens incredible new doors for the medical community. Researchers who engineer specific bacteria for large-scale manufacturing, study exactly how different gut microbes interact with the human body, or actively develop brand-new antibiotics, can now use this fresh data to easily choose an appropriate retro-based editing system for whatever species they work with. Shipman hopes this highly effective editing technology will continue to spread rapidly across the globe, allowing scientists to uncover brand new biology and directly intervene in deadly diseases.